Diagnostics based on nucleic acid sequence variant profiling: PCR, hybridization, and NGS approaches - ScienceDirect

Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing | Nature Communications

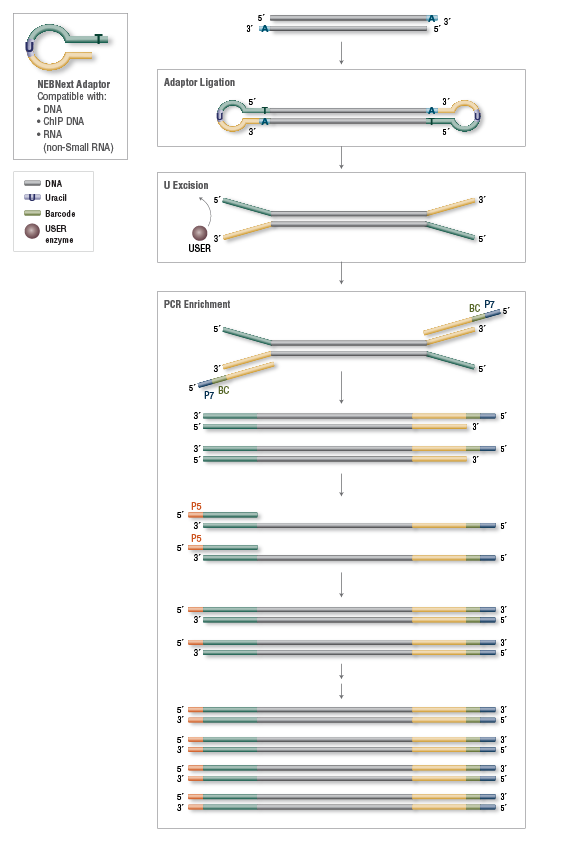
Overview of the different methods used for adapter addition to antibody... | Download Scientific Diagram
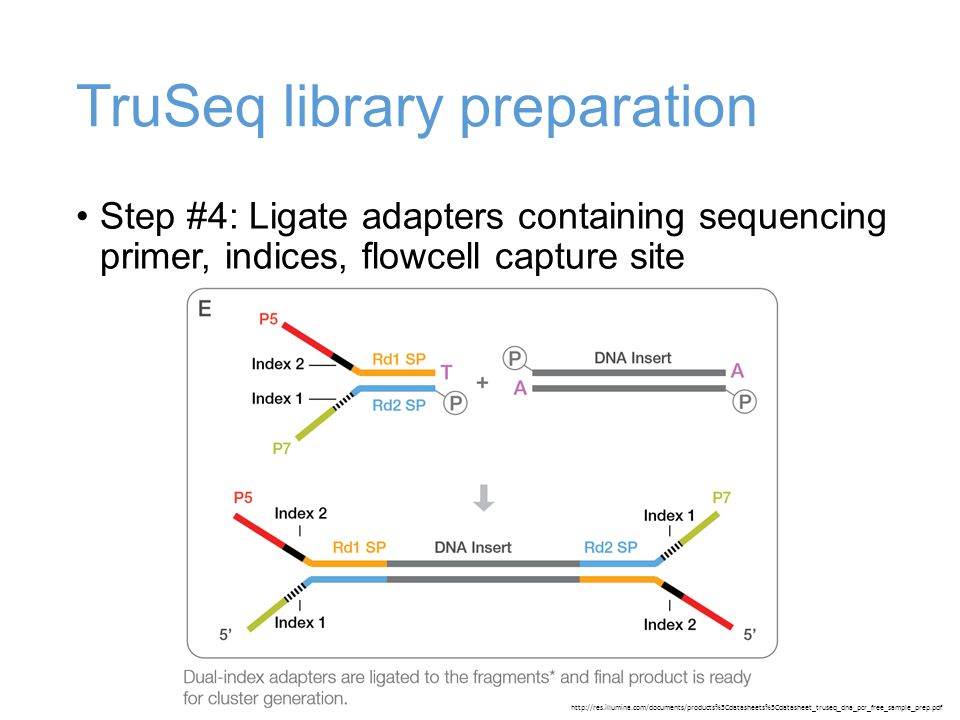
Quantitative Bias in Illumina TruSeq and a Novel Post Amplification Barcoding Strategy for Multiplexed DNA and Small RNA Deep Sequencing | PLOS ONE

Incorporation of unique molecular identifiers (UMIs) in TruSeq adapters improves the accuracy of quantitative sequencing | RNA-Seq Blog

Incorporation of unique molecular identifiers (UMIs) in TruSeq adapters improves the accuracy of quantitative sequencing | RNA-Seq Blog
Indexing and Barcoding for Illumina NextGen Sequencing (ELK 10/2011) There are 2 main strategies for indexing/barcoding (which a

Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge

Figure 1 from Quantitative Bias in Illumina TruSeq and a Novel Post Amplification Barcoding Strategy for Multiplexed DNA and Small RNA Deep Sequencing | Semantic Scholar
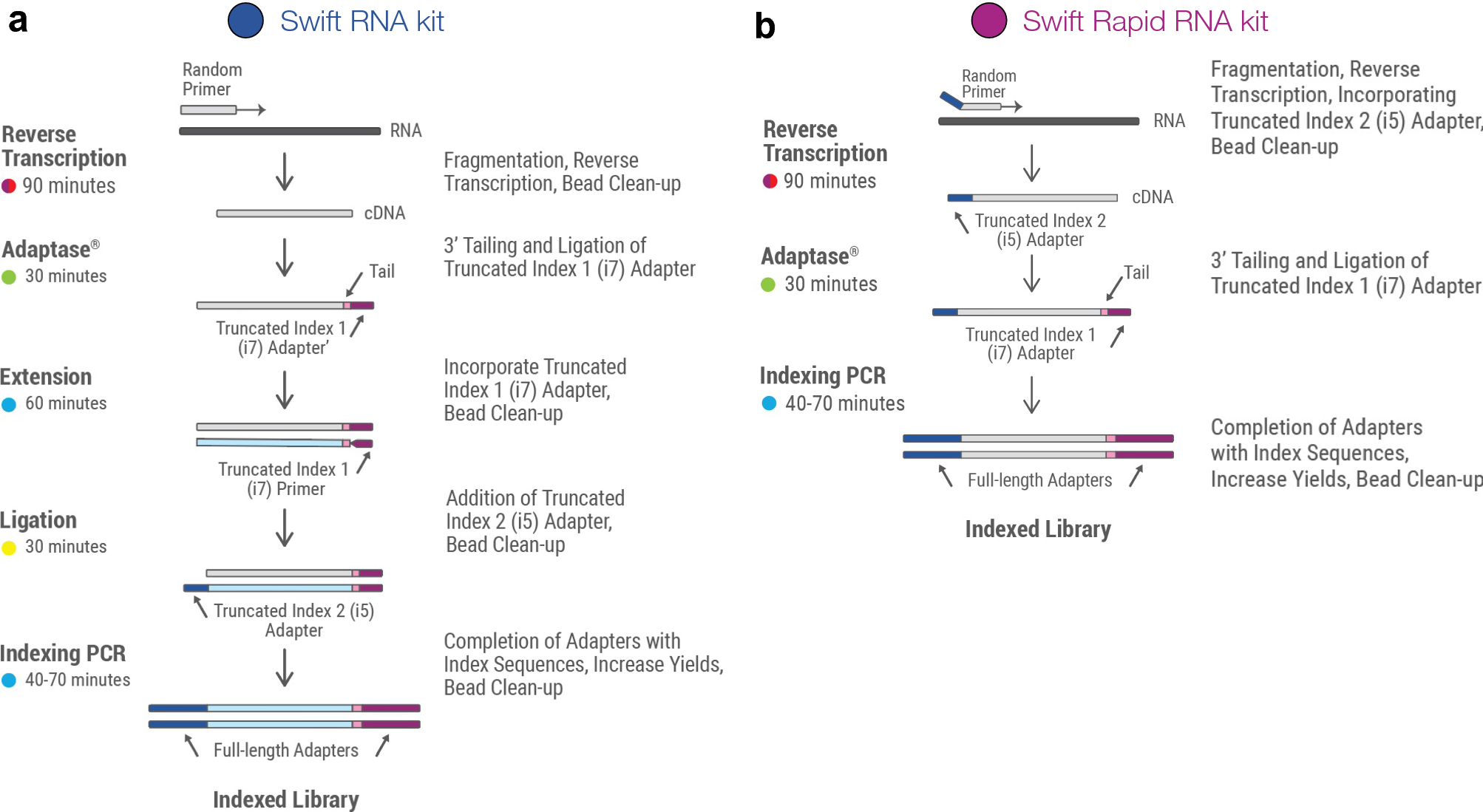
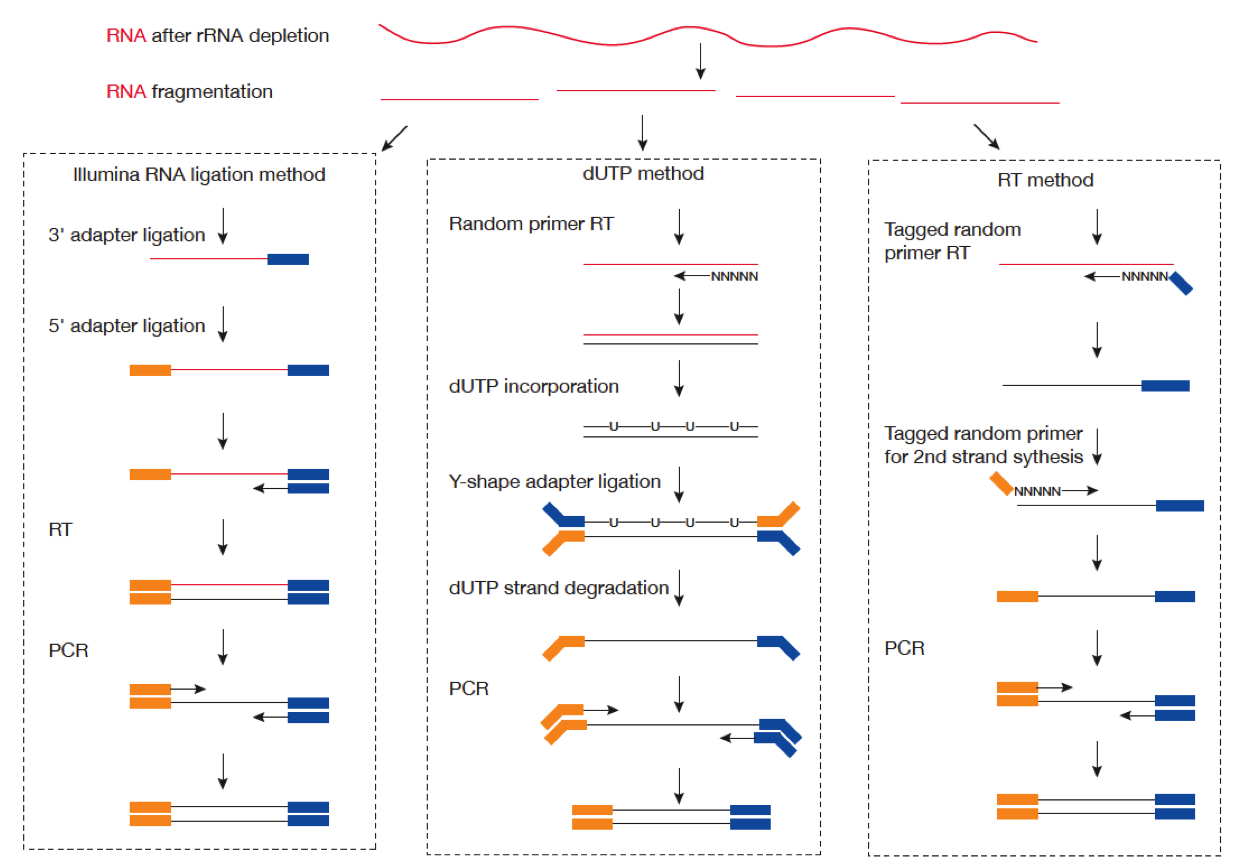
Systematic comparative analysis of strand-specific RNA-seq library preparation methods for low input samples | Scientific Reports
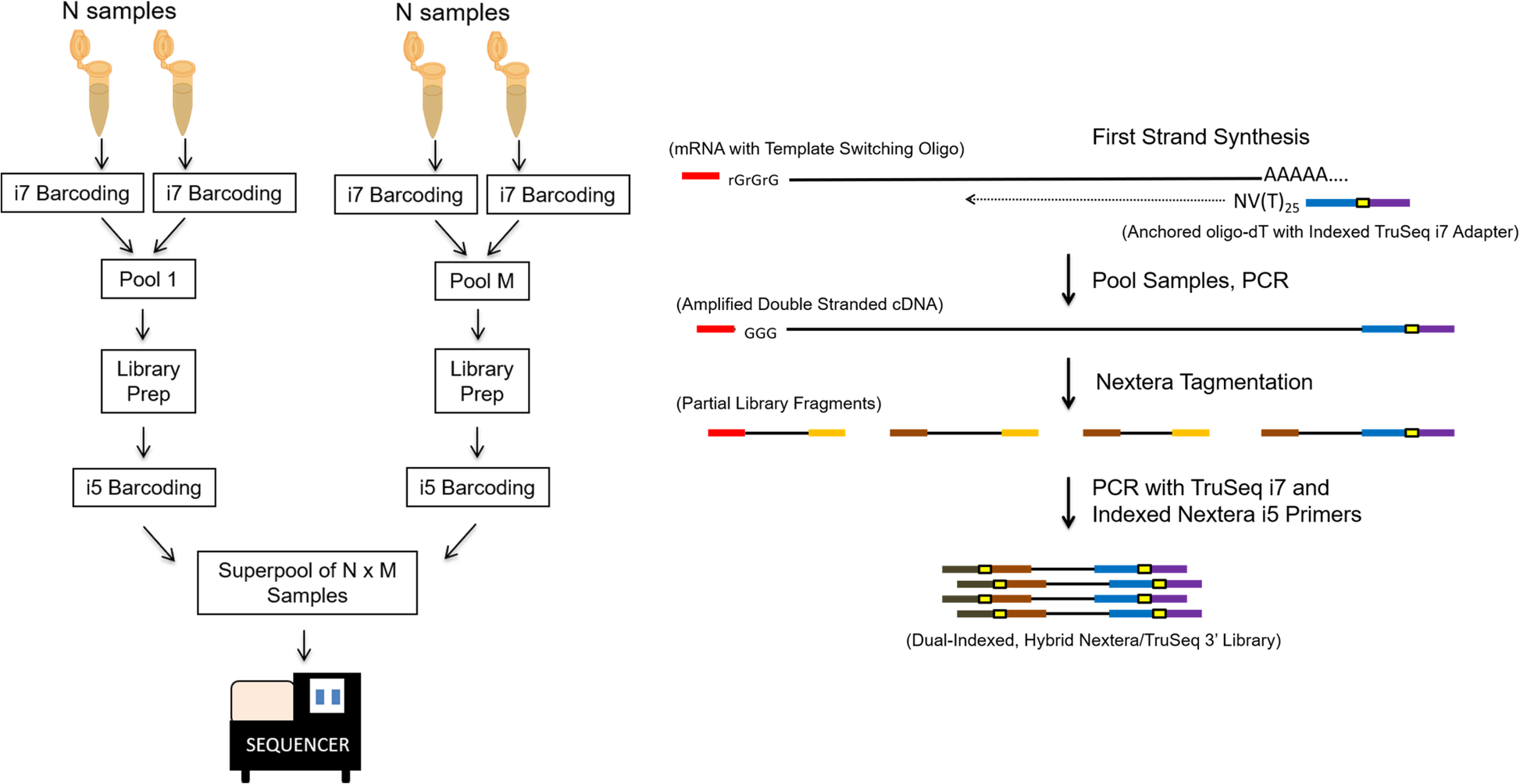
3'Pool-seq: an optimized cost-efficient and scalable method of whole-transcriptome gene expression profiling | BMC Genomics | Full Text