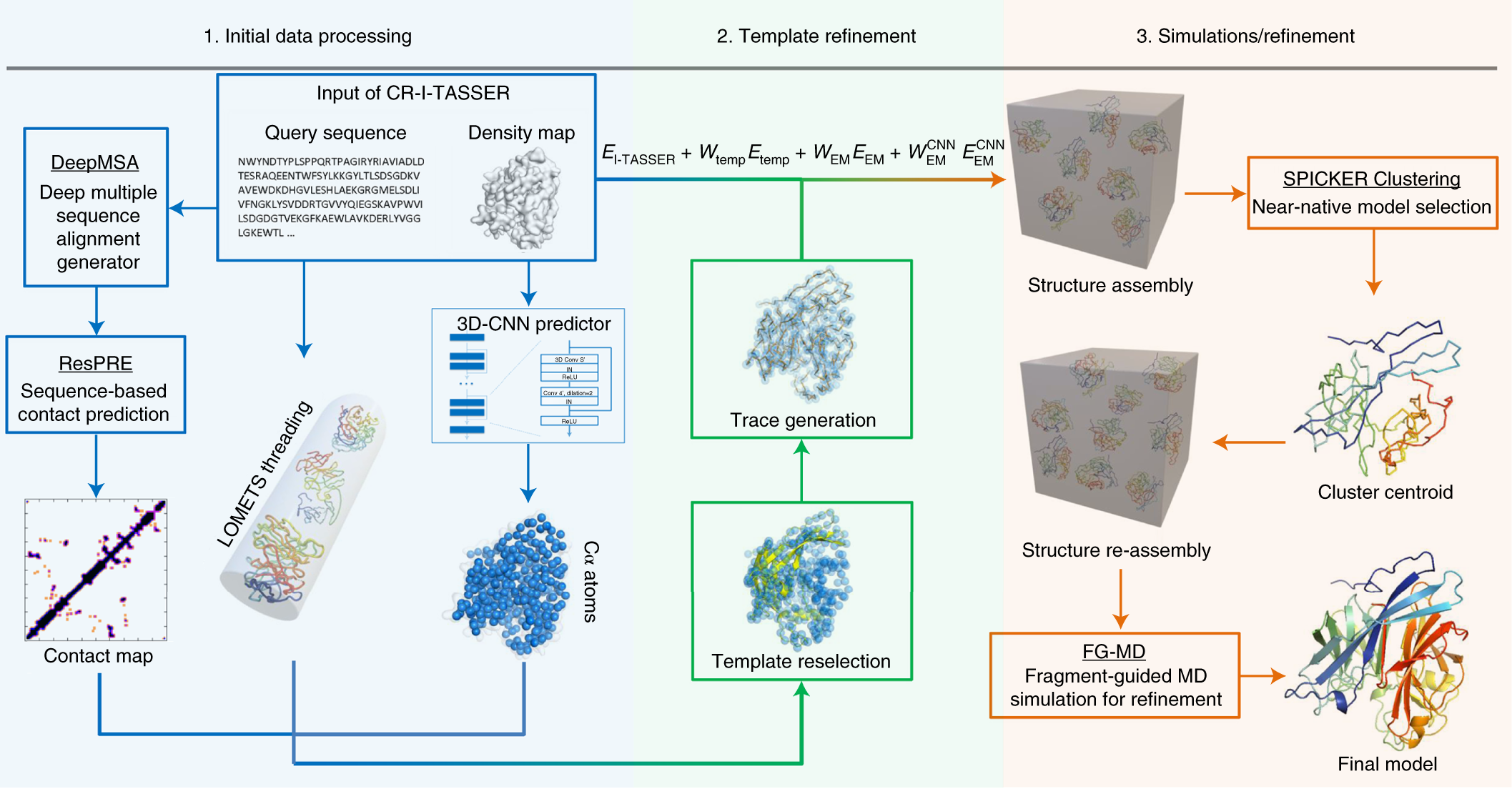
CR-I-TASSER: assemble protein structures from cryo-EM density maps using deep convolutional neural networks | Nature Methods

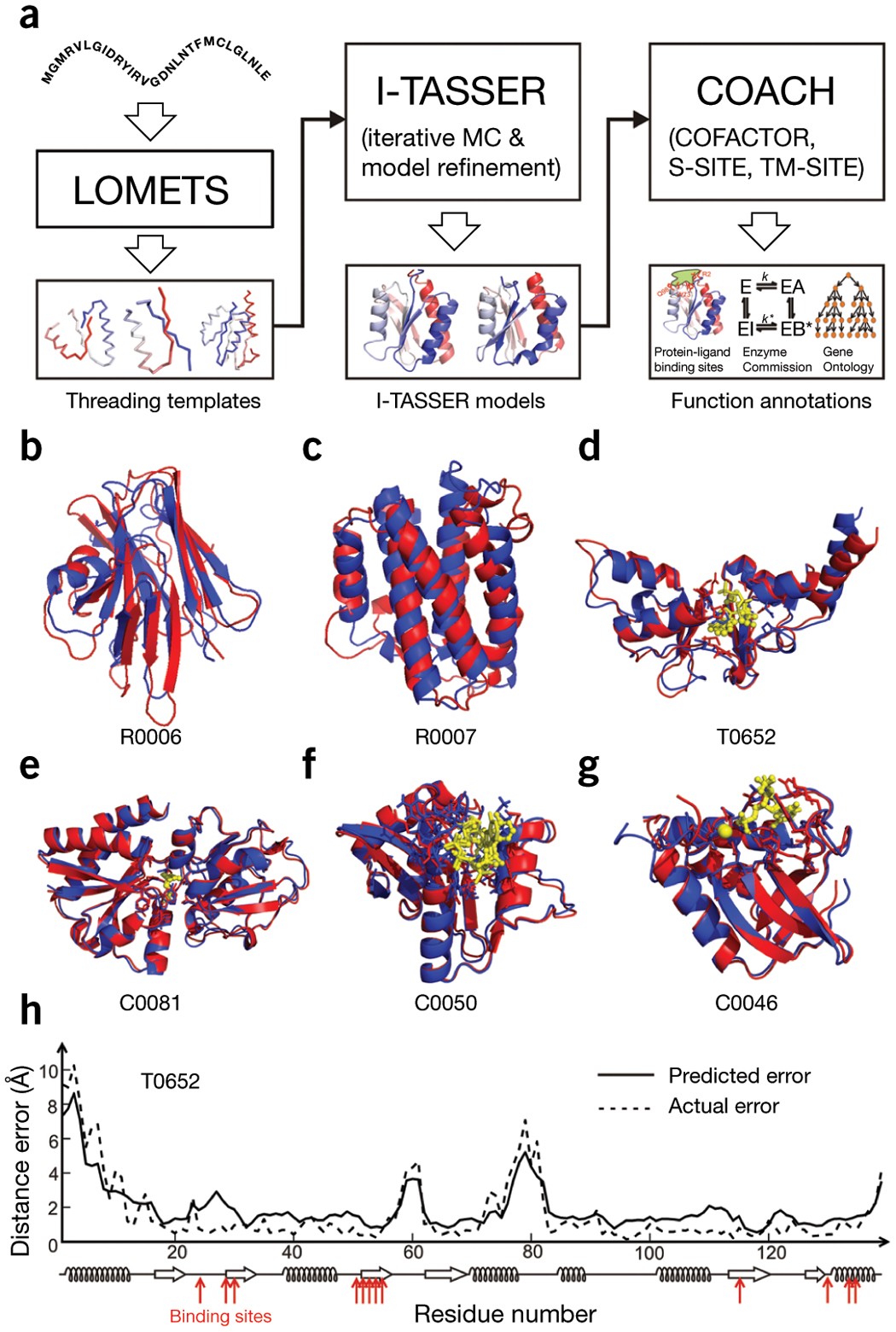
Ultrafast end-to-end protein structure prediction enables high-throughput exploration of uncharacterized proteins | PNAS
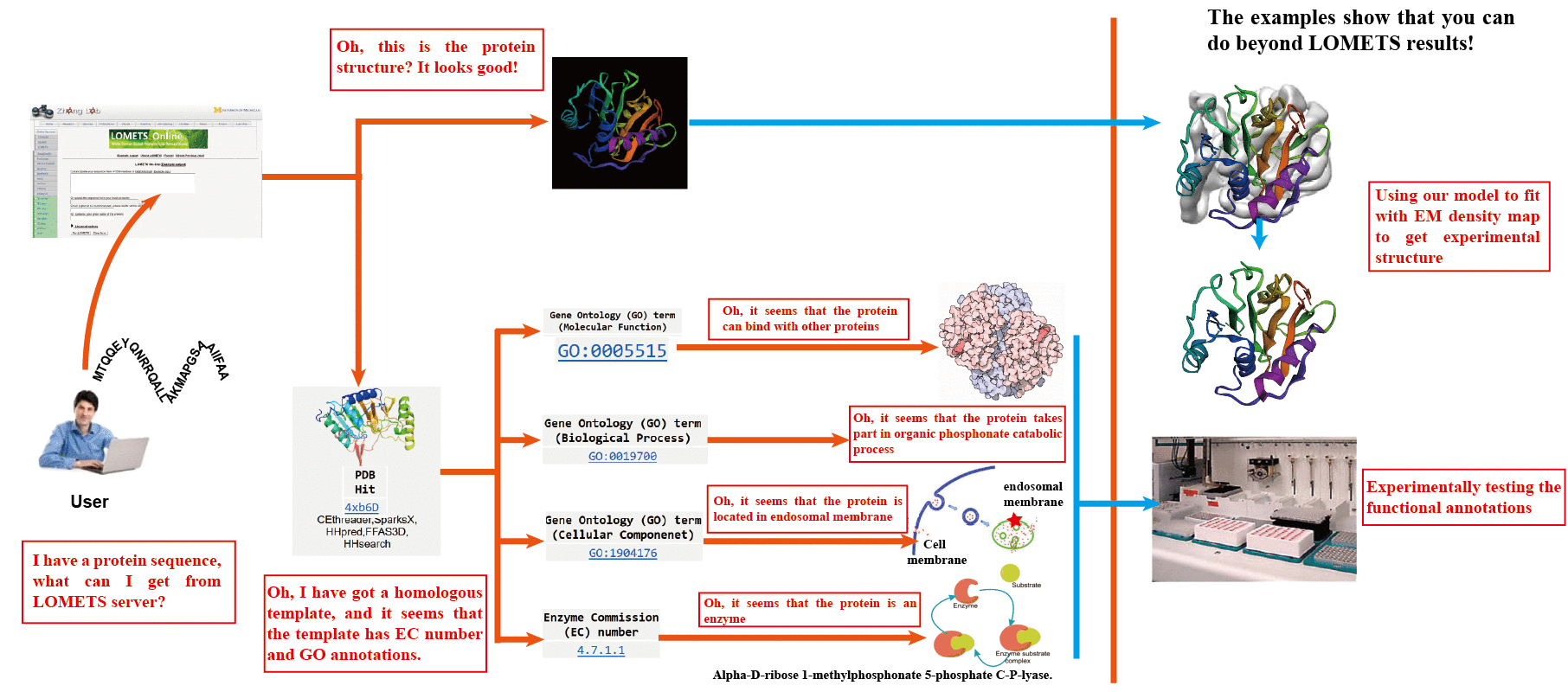
![PDF] I-TASSER gateway: A protein structure and function prediction server powered by XSEDE | Semantic Scholar PDF] I-TASSER gateway: A protein structure and function prediction server powered by XSEDE | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6799e492f2e6405f20010a157973295fa8ac2b64/7-Figure5-1.png)
PDF] I-TASSER gateway: A protein structure and function prediction server powered by XSEDE | Semantic Scholar
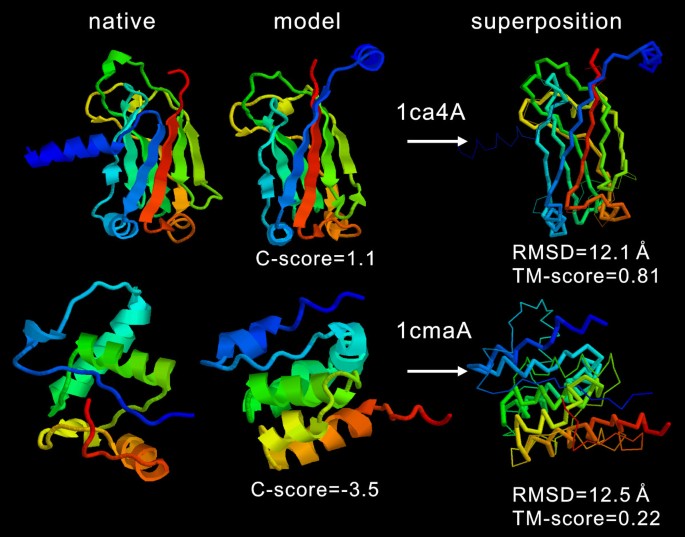
![PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/682fc6a2dcf6edaadacb1a2f90fdd137aef28d15/2-Figure1-1.png)
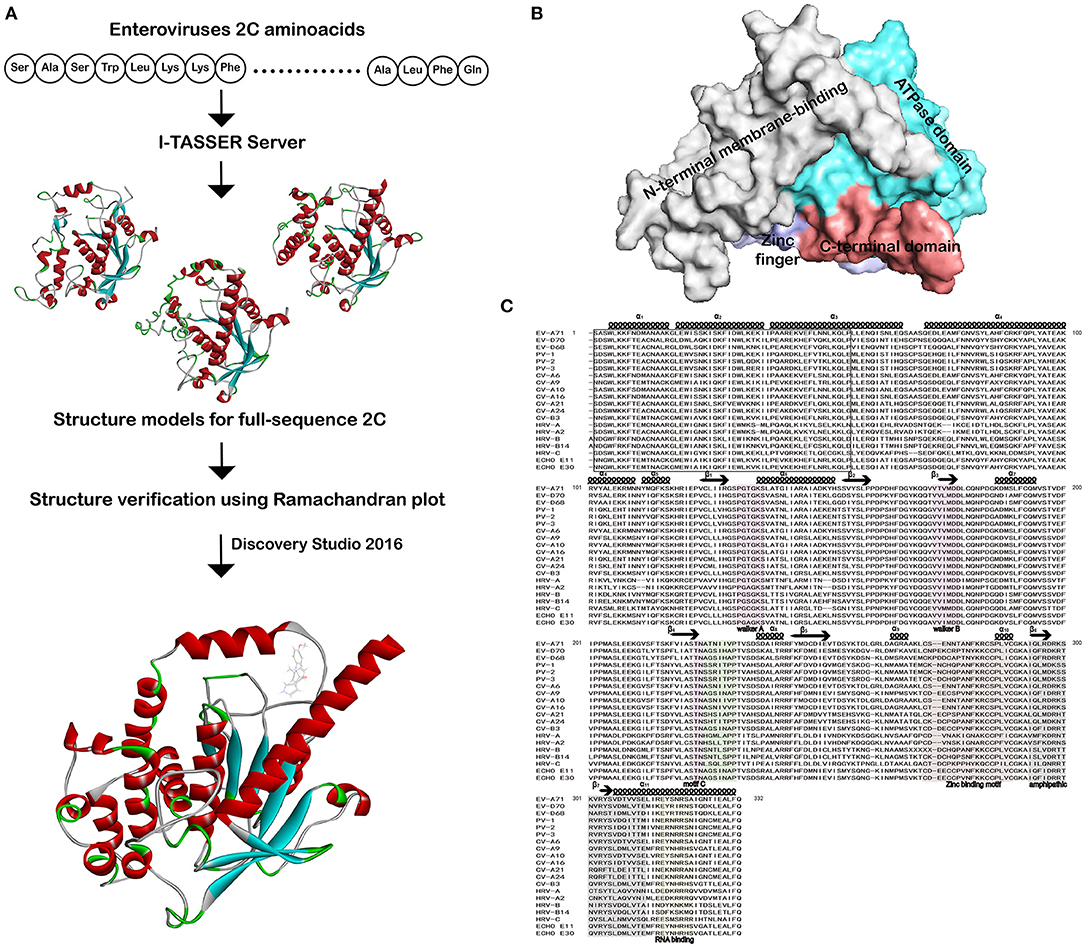
PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar

I-TASSER-MTD: a deep-learning-based platform for multi-domain protein structure and function prediction | Nature Protocols

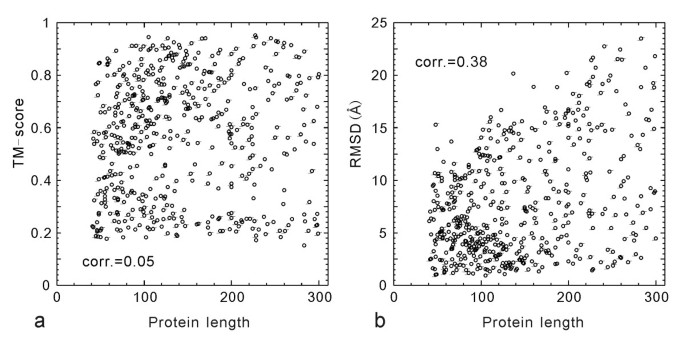
Template‐based modeling and free modeling by I‐TASSER in CASP7 - Zhang - 2007 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library
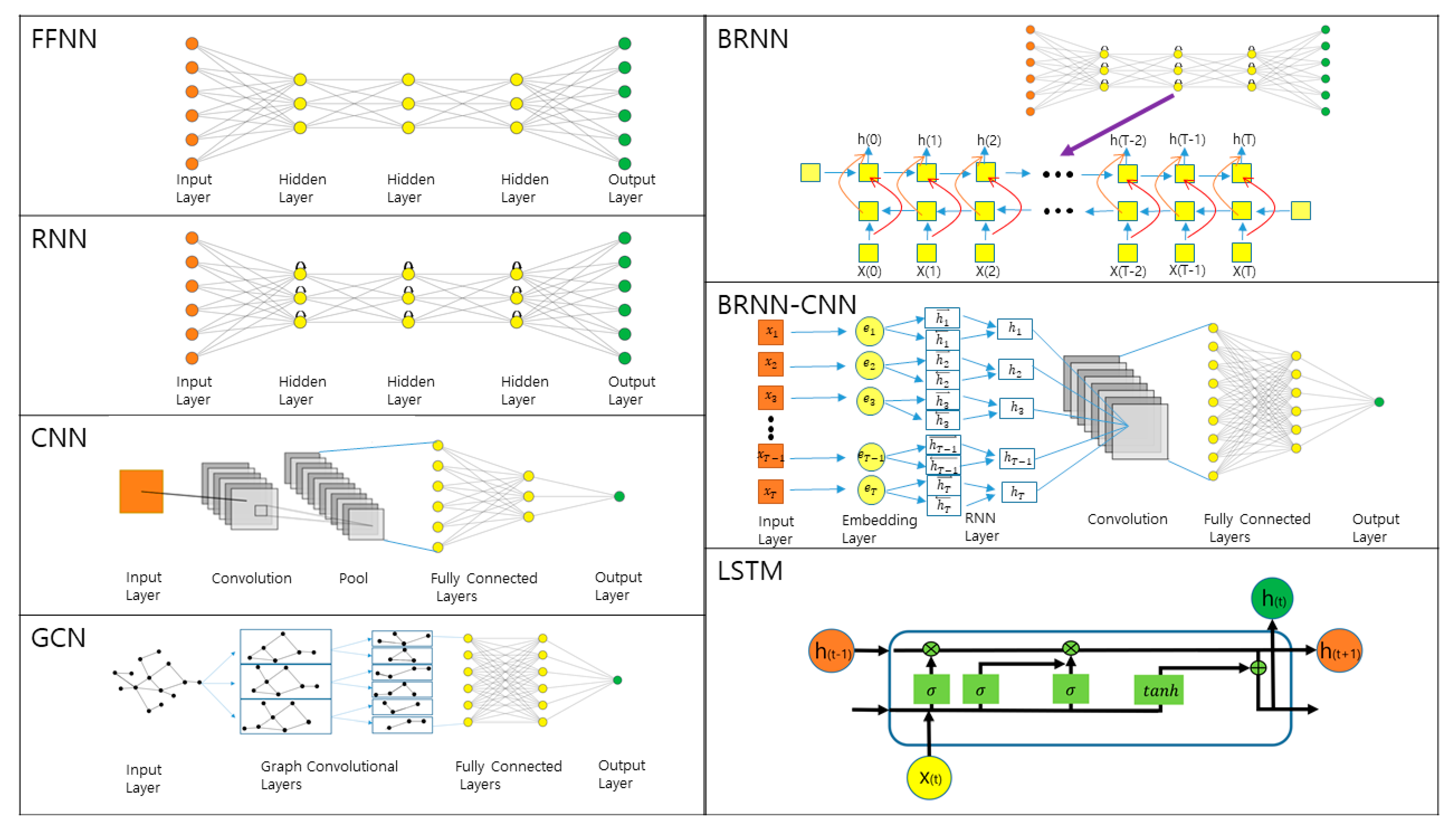
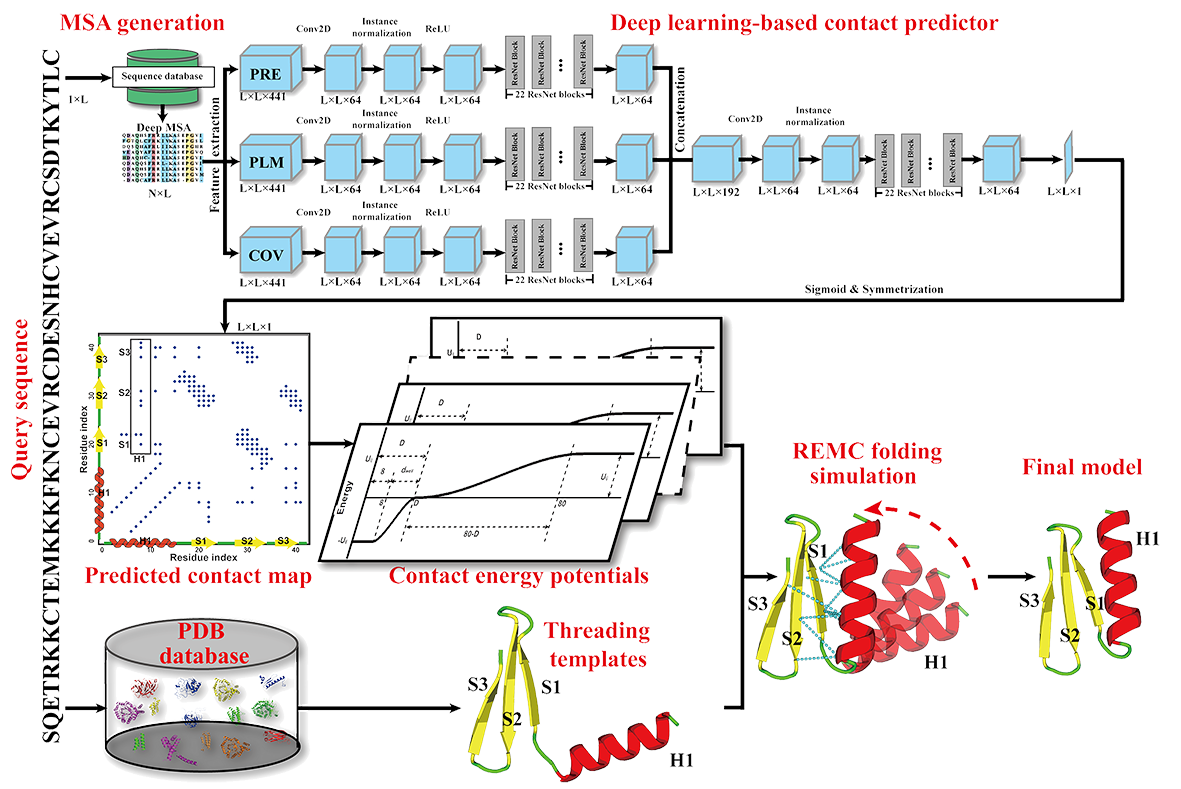
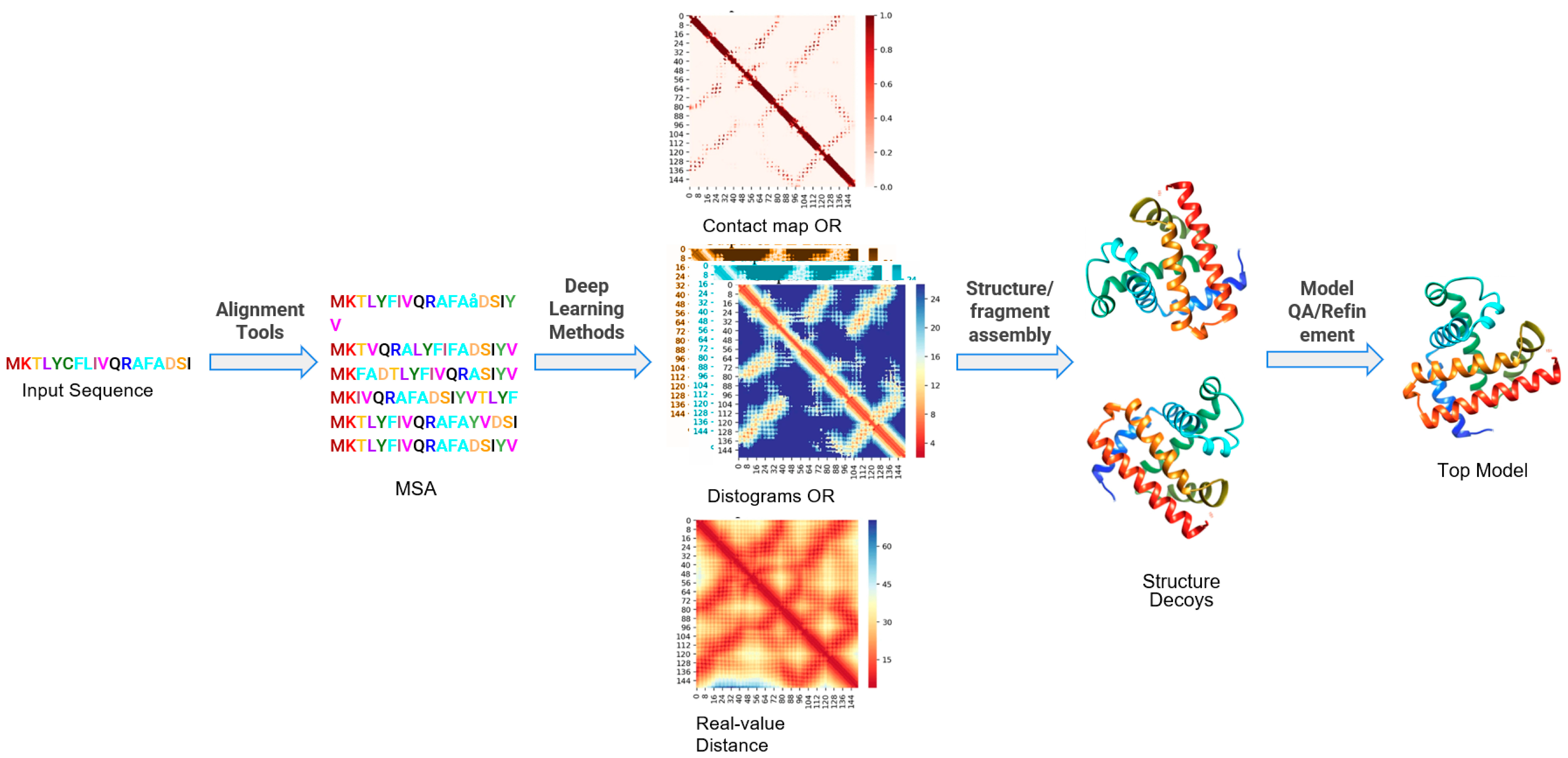
IJMS | Free Full-Text | Recent Applications of Deep Learning Methods on Evolution- and Contact-Based Protein Structure Prediction
![PDF] I-TASSER gateway: A protein structure and function prediction server powered by XSEDE | Semantic Scholar PDF] I-TASSER gateway: A protein structure and function prediction server powered by XSEDE | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6799e492f2e6405f20010a157973295fa8ac2b64/9-Figure7-1.png)

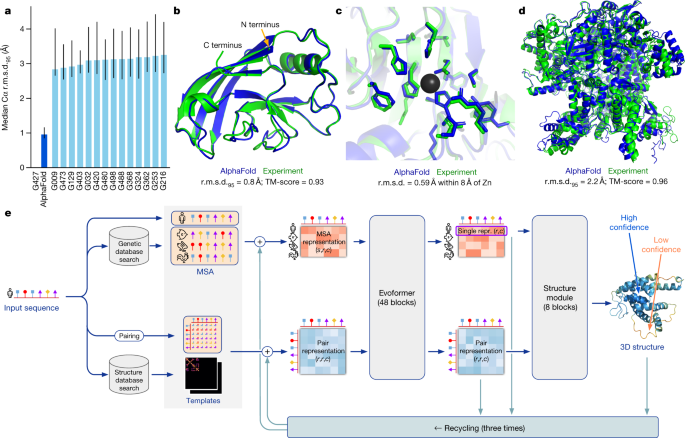
![General workflow of I-TASSER for protein structure prediction [30] | Download Scientific Diagram General workflow of I-TASSER for protein structure prediction [30] | Download Scientific Diagram](https://www.researchgate.net/publication/281540998/figure/fig1/AS:281366399864834@1444094385414/General-workflow-of-I-TASSER-for-protein-structure-prediction-30.png)